Case Study

Iddo Weiner
Zero-Shot Design of a Biobetter Cetuximab: Enhancing EGFR Binding with ConvergeAB™
TL;DR ConvergeAB™ was used in a zero-shot setting to design a biobetter version of cetuximab targeting EGFR. The lead Converge-designed antibody demonstrated consistently superior binding kinetics, with ~2.1× higher affinity than cetuximab and ~4.4× higher than a computationally designed competitor and includes 6 total edits, 2 in the CDR sections, all while matching or exceeding the developability profile of the original cetuximab. Read the full Forbes cover
Introduction
Epidermal Growth Factor Receptor (EGFR) is a transmembrane tyrosine kinase that plays a central role in regulating cell proliferation, survival, and differentiation. Upon binding of its natural ligands, such as epidermal growth factor (EGF), EGFR undergoes dimerization and autophosphorylation, triggering downstream signaling cascades. Dysregulation of EGFR, through overexpression or mutation, is a well-established driver of oncogenesis. Elevated EGFR activity has been implicated across multiple solid tumors, most notably in colorectal cancer, head and neck squamous cell carcinoma (HNSCC), and non-small cell lung cancer (NSCLC). In these contexts, aberrant EGFR signaling promotes uncontrolled cell growth, resistance to apoptosis, and enhanced metastatic potential, making it a compelling therapeutic target.
Cetuximab is a chimeric monoclonal antibody designed to target the extracellular domain of EGFR. By binding to the receptor with high specificity, cetuximab sterically blocks ligand interaction, thereby preventing receptor activation and downstream signaling. Clinically, cetuximab has become a cornerstone therapy in EGFR-driven malignancies, particularly in metastatic colorectal cancer and recurrent or metastatic HNSCC. It is widely used in combination with chemotherapy or radiation, with global adoption supported by regulatory approvals across the world.
At the molecular level, the therapeutic efficacy of cetuximab is fundamentally governed by its competition with endogenous ligands such as EGF for binding to EGFR. This competitive interaction is dictated by binding affinity, commonly quantified by the dissociation constant (KD). A lower KD reflects tighter binding and greater competitive advantage
. In tumors where ligand concentrations are high, cetuximab must effectively outcompete EGF to suppress receptor activation. Insufficient binding affinity can allow residual signaling, contributing to suboptimal response and disease progression. Therefore, optimizing KD is not merely a biochemical consideration but a critical determinant of clinical efficacy, influencing both depth and durability of response in patients.
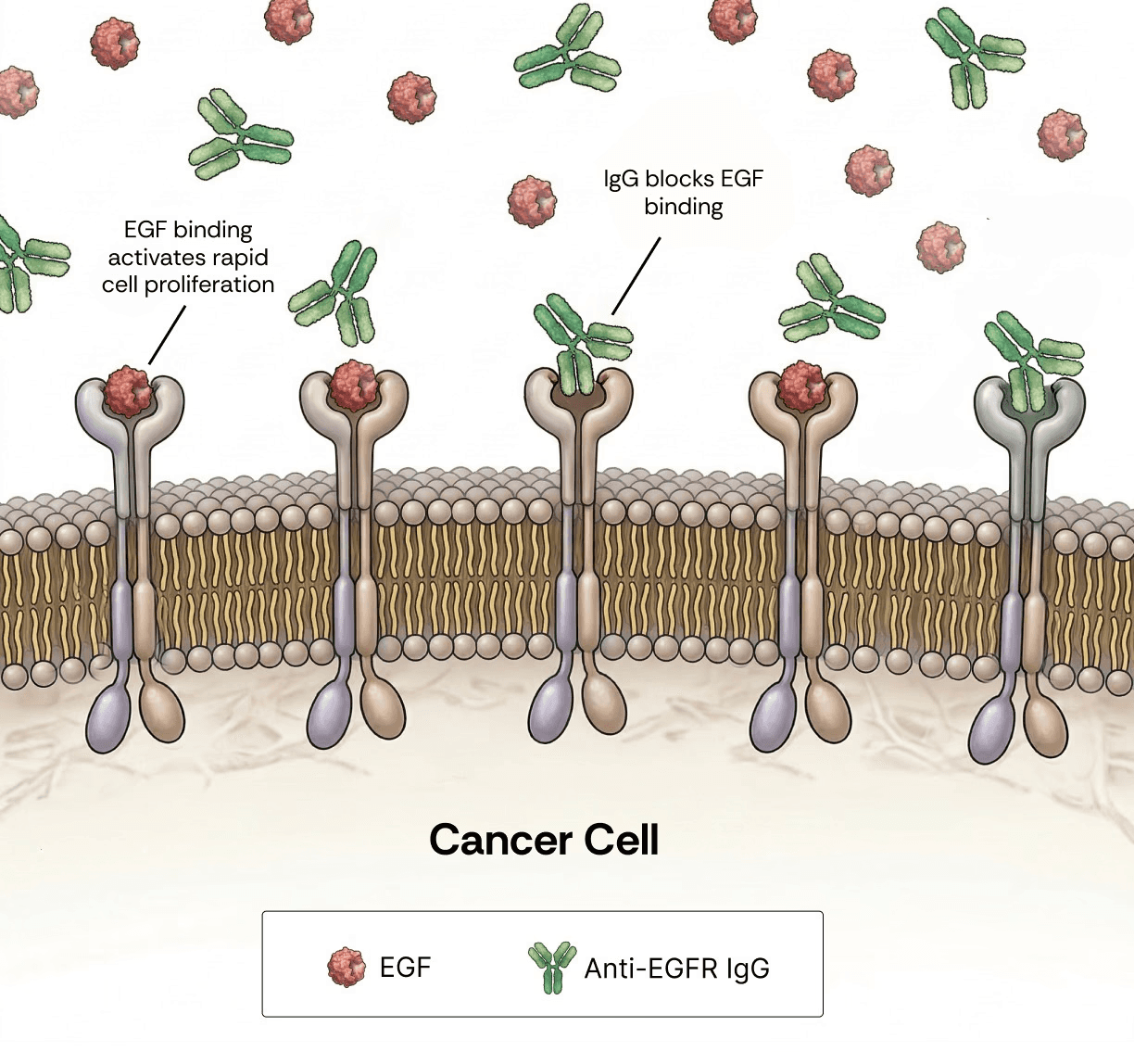
Figure 1. Competitive binding at EGFR is a key determinant of the therapeutic efficacy of anti-EGFR antibodies. Epidermal growth factor (EGF, red) and anti-EGFR monoclonal IgG (green) compete for binding to the EGFR transmembrane receptor on the surface of cancer cells. Engagement of EGFR by EGF activates proliferative signaling pathways that drive tumor growth and progression. In contrast, antibody binding blocks ligand-induced receptor activation, leading to growth inhibition, cell cycle arrest, or apoptosis. The outcome of this competition is governed by binding kinetics, with lower KD antibodies exhibiting superior competitive advantage over endogenous ligands.
Results
To generate a biobetter version of cetuximab, we prompted ConvergeAB™ with the original cetuximab sequence together with the EGFR target sequence. No task-specific or ad hoc training was done prior to the initiation of this affinity maturation campaign, effectively creating a zero shot effort. In brief, ConvergeAB™ is a multi-step platform in which sequence diversity is introduced into a seed antibody using a proprietary protein language model, generating ~100,000 target-aware candidates. These candidates are subsequently filtered through a series of explicit and orthogonal predictors that evaluate key properties, including binding affinity, structural compatibility (docking), expressibility, thermal stability, and solubility. Additional details on the ConvergeAB™ platform can be found here: https://converge-bio.com/solutions/converge-ab.
The top 10 predicted candidates of the campaign were selected for experimental validation. Each variant was expressed and screened for binding using single-concentration surface plasmon resonance (SPR). The highest-performing candidate from this screen was advanced for full kinetic characterization, including precise KD determination. As benchmarks, we included both the original cetuximab and a recently published anti-EGFR antibody from Cradle Bio, reported as a top performer in a protein design competition (https://www.adaptyvbio.com/blog/po104/).
All three antibodies (Converge, Cradle, and cetuximab) were produced in the IgG1 format, consistent with the clinical form of cetuximab. For rigorous head-to-head comparison, the three antibodies were analyzed side-by-side in two unique SPR repeats (#1 and #2) using a six-analyte, multi-cycle kinetic SPR assay. Across all measured parameters (kon, koff, and KD), and in both biological repeats, the Converge-designed antibody consistently demonstrated superior performance, achieving an average binding affinity 2.1-fold stronger than cetuximab and 4.4-fold stronger than the Cradle-derived antibody.
Antibody | SPR repeat | Kon (1/Ms) | Koff (1/s) | KD (pM) |
Converge | #1 | 5.76x106 | 1.28x10-3 | 222 |
#2 | 3.1x106 | 1.27x10-3 | 409 | |
Mean | 4.43x106 | 1.275x10-3 | 315 | |
Cetuximab | #1 | 3.23x106 | 1.81x10-3 | 560 |
#2 | 2.25x106 | 1.77x10-3 | 786 | |
Mean | 2.74x106 | 1.79x10-3 | 673 | |
Cradle | #1 | 3.92x106 | 4.32x10-3 | 1,100 |
#2 | 2.53x106 | 4.21x10-3 | 1,660 | |
Mean | 3.22x106 | 4.265x10-3 | 1,380 |
Table 1. Summary of SPR kinetic parameters for anti-EGFR antibodies. Three anti-EGFR antibodies were evaluated side-by-side for binding to EGFR. Each SPR repeat represents an independent, parallel measurement of all three antibodies under identical conditions. The association rate constant (kon) reflects the rate of antibody–antigen complex formation, with higher values indicating faster binding. The dissociation rate constant (koff) reflects the rate of complex dissociation, with lower values indicating greater complex stability. The equilibrium dissociation constant (KD), calculated as koff / kon, summarizes overall binding affinity, with lower values corresponding to stronger binding.
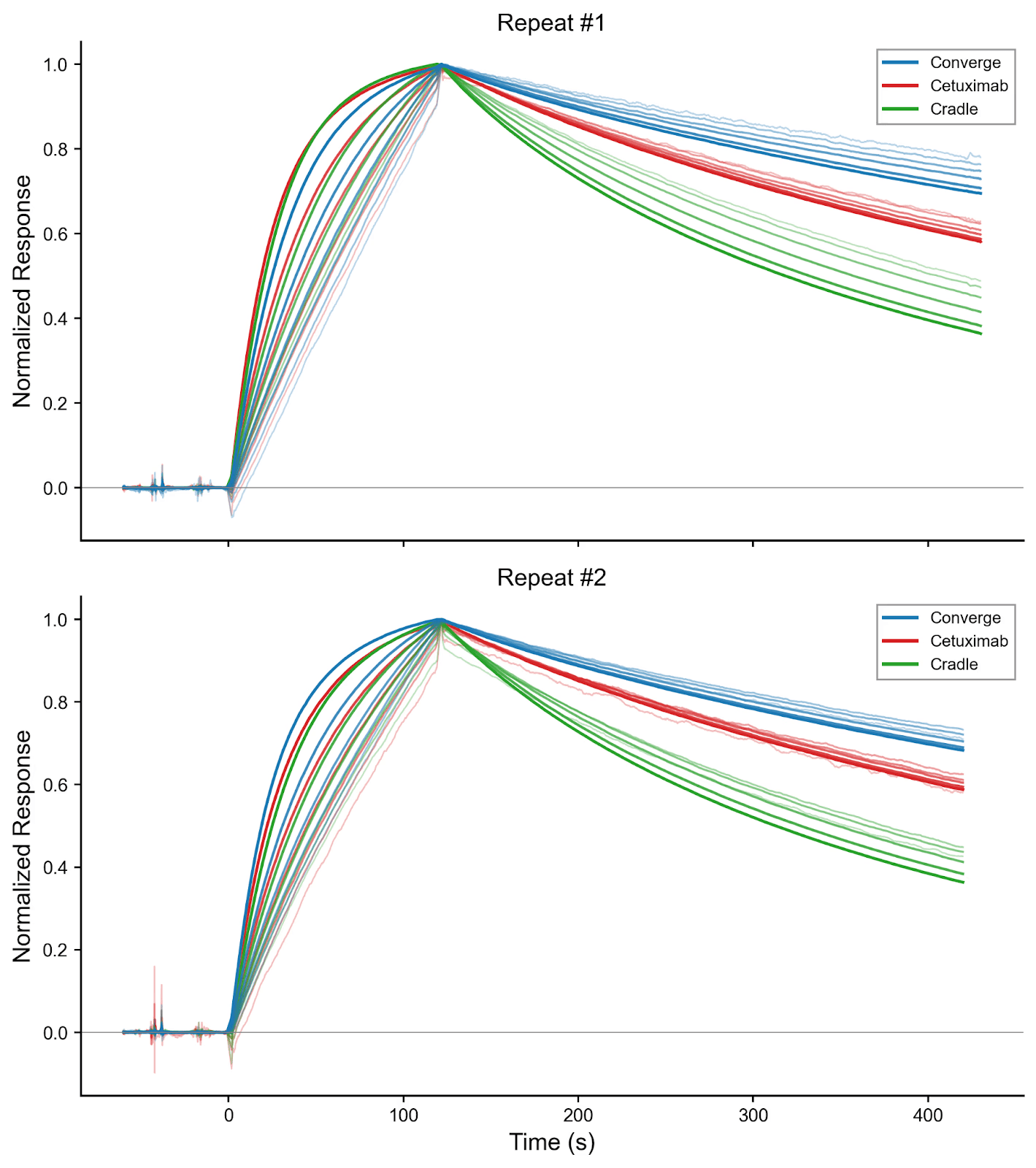
Figure 2. SPR multikinetic raw data of both repeats. SPR sensorgrams from each repeat were normalized to enable side-by-side comparison. EGFR was used as the analyte at the following concentrations (nM): 0.2, 0.39, 0.78, 1.6, 3.1, 6.2, and 12.5. Curve thickness corresponds to analyte concentration, with the boldest curve representing the highest concentration.
Sequence analysis reveals the Converge sequence had a total of six edits compared to cetuximab, comprising four substitutions to alanine (VH T61A, VH S87A, VL V9A, and VL N93A) and two asparagine-to-aspartate substitutions (VH N88D and VL N32D), distributed across both framework and CDR regions. To assess whether these affinity gains were associated with large-scale structural remodeling, we predicted the 3D structures of both antibodies and aligned them. The resulting overall RMSD was very low, at 0.16 Å, indicating that the Converge variant preserves the global cetuximab binding architecture. This is an important observation, as it argues against a wholesale change in fold or binding mode and instead supports a mechanism in which affinity improvement is driven by highly localized structural and energetic optimization within an otherwise conserved paratope.
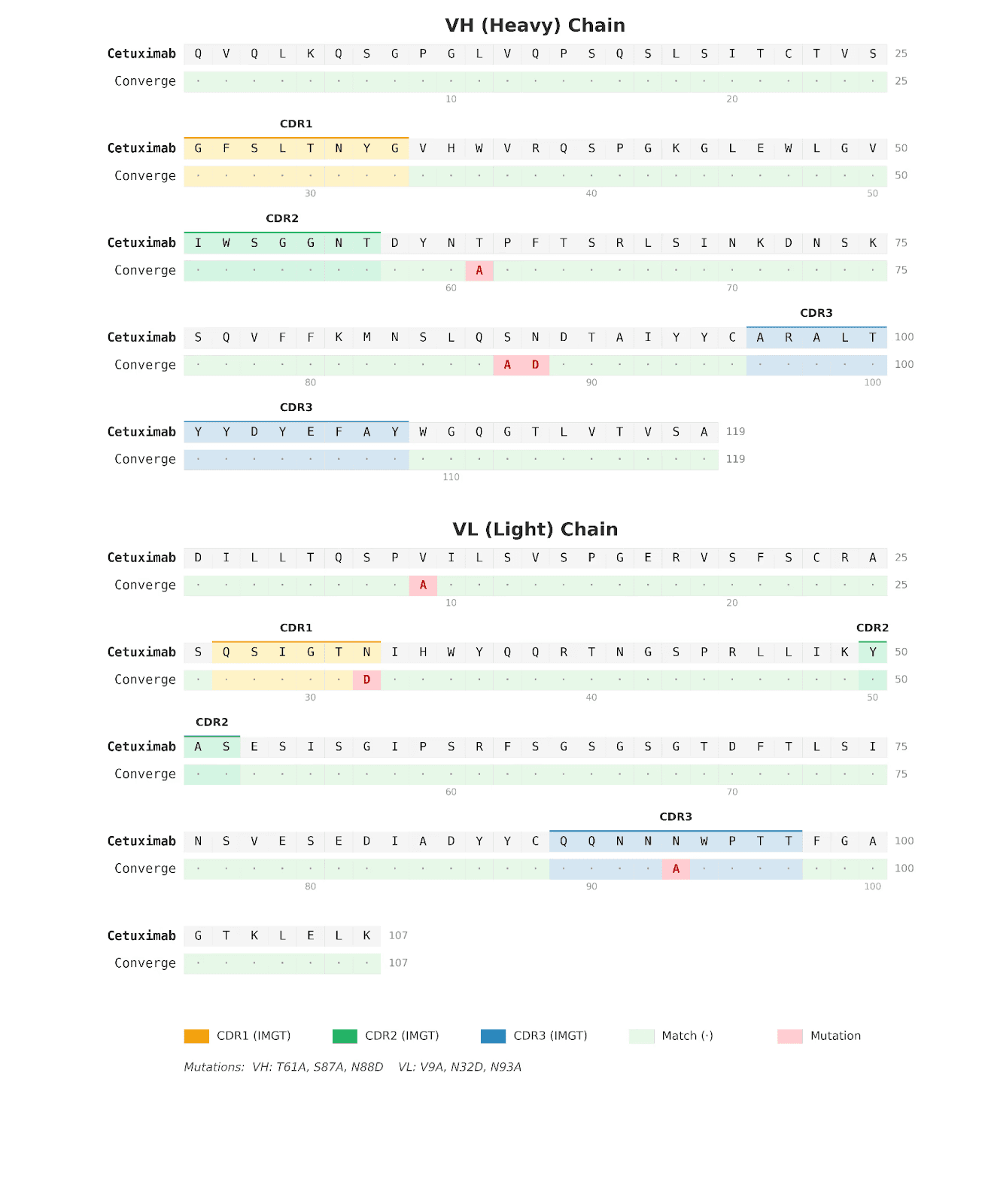
Figure 3. Sequence alignment of cetuximab and Converge-designed anti-EGFR antibody. Amino acid sequence alignment comparing cetuximab with the Converge-derived anti-EGFR IgG marks the positions of the six edits introduced relative to cetuximab.
This interpretation is consistent with the nature of the edits. The alanine substitutions are expected to reduce side-chain bulk and remove potentially suboptimal polar functionality, which can improve local packing, decrease conformational heterogeneity, and favor a more pre-organized binding-competent state. Likewise, the Asparagine to Aspartate substitutions introduce negative charge and altered hydrogen-bonding geometry, which may strengthen electrostatic complementarity with EGFR or stabilize productive local conformations. The structural comparison further supports this localized mechanism: although the global RMSD remains minimal, the N93A substitution produces a 0.66 Å shift at the Ca position and displaces Trp94 by 0.5 Å. These are small but meaningful local rearrangements, especially in an antibody paratope, where sub-angstrom movements can measurably alter side-chain presentation, loop packing, and epitope complementarity without changing the overall fold.
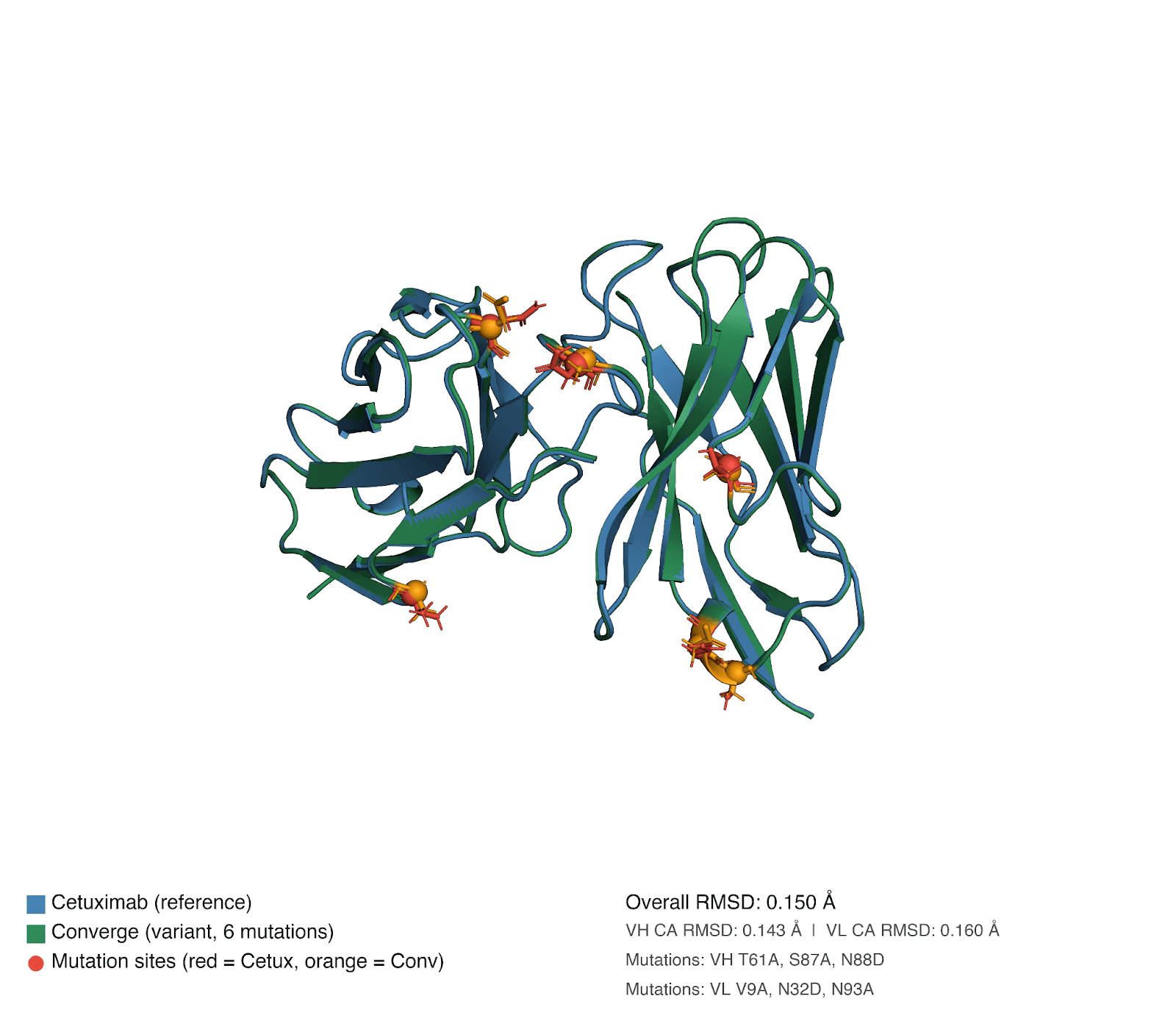
Figure 4. Structural alignment of cetuximab and Converge-designed anti-EGFR antibody. Structural superposition of cetuximab and the Converge-derived anti-EGFR IgG highlights overall architectural similarity.
Taken together, the sequence and structure analyses support a compelling model for the experimentally confirmed affinity improvement. Rather than arising from major structural divergence from cetuximab, the Converge sequence appears to retain the parental architecture while introducing a set of cooperative micro-adjustments that fine-tune the binding interface. The very low overall RMSD, combined with specific local shifts associated with N93A and neighboring Trp94, is therefore fully consistent with an affinity maturation mechanism based on paratope preorganization, improved local packing, and targeted electrostatic optimization.
Developability Profile
Affinity gains are clinically valuable only when accompanied by acceptable biophysical behavior. To assess whether the localized sequence optimizations preserved cetuximab's drug-like properties, we evaluated Converge, cetuximab, and Cradle against an orthogonal panel of developability assays: hydrophobic interaction chromatography (HIC), dynamic light scattering (DLS), differential scanning fluorimetry (DSF), and a polyspecificity reagent (PSR) ELISA panel. Across all four parameters, the Converge antibody showed a profile comparable to or better than cetuximab, presenting no developability red flags.
Hydrophobicity (HIC): All three antibodies eluted as predominantly single, sharp peaks (>95% main-peak area at 214 nm), indicating no aggregation or fragmentation liabilities. The Converge variant displayed modestly higher hydrophobicity than cetuximab (RRT of 1.25 vs. 1.08), which is consistent with the introduction of four alanine substitutions. However, this profile remains well within the developable envelope expected for clinical-stage IgG1s.
Antibody | RT (min) | RRT (vs Trastuzumab) | Main peak Area% (214 nm) |
Converge | 8.108 | 1.253 | 98.21 |
Cetuximab | 6.967 | 1.076 | 95.94 |
Cradle | 8.780 | 1.356 | 99.23 |
Trastuzumab (control) | 6.473 | 1.000 | — |
Table 2. HIC retention times for the three antibodies. RT, retention time at the main 214 nm peak; RRT, relative retention time vs. trastuzumab control (RT = 6.473 min). Higher RRT indicates greater surface hydrophobicity. Main-peak Area% (214 nm) reports purity/monomer-dominance of the chromatogram.
Hydrodynamic Size and Dispersity (DLS): The Converge candidate demonstrated the most compact and uniform structural profile among the tested antibodies, with the smallest hydrodynamic radius (5.85 nm) and the lowest polydispersity (3.95% PD). These metrics fall strictly within the benchmark envelope, confirming a highly monodisperse, monomeric population.
Antibody | Radius (nm) | Diameter (nm) | %PD | Norm. intensity (Cnt/s) |
Converge | 5.85 | 11.65 | 3.95 | 1.12 × 10⁷ |
Cetuximab | 6.80 | 13.55 | 10.45 | 1.92 × 10⁷ |
Cradle | 7.40 | 14.75 | 10.05 | 4.52 × 10⁷ |
Trastuzumab (control) | 5.70 | 11.35 | 5.45 | 1.32 × 10⁷ |
Table 3. DLS results (mean of two replicates). Lower hydrodynamic radius and lower %PD reflect a more compact, monodisperse population. Normalized intensity is reported for completeness but is sensitive to concentration and is not a developability metric on its own.
Thermal Stability (DSF): The thermal unfolding profile (Tonset, Tm1, Tm2) confirmed that all variants maintained adequate thermal stability. The Converge variant's Tonset (59.54 °C) and Tm1 (68.34 °C) were marginally lower than cetuximab's (~1.2 °C), an expected minor destabilization from the alanine edits. Importantly, Converge’s Tm2 (80.96 °C) was essentially equivalent to cetuximab's (82.72 °C), indicating that the core of the Fab domain remains well folded.
Antibody | T_onset (°C) | Tm1 (°C) | Tm2 (°C) |
Converge | 59.54 | 68.34 | 80.96 |
Cetuximab | 60.89 | 69.49 | 82.72 |
Cradle | 61.62 | 68.86 | 77.48 |
Trastuzumab (control) | 62.81 | 70.15 | 80.77 |
Table 4. DSF transition temperatures. Tonset is the temperature at which unfolding begins; Tm1, Tm2 are successive thermal transitions, typically attributed to CH2, CH3/Fab, and Fab core unfolding events in human IgG1.
Polyspecificity (PSR ELISA): Tested across three orthogonal substrates (bovine viral particle, dsDNA, and insulin), both Converge and cetuximab proved essentially indistinguishable from a clinical negative-control antibody (Infliximab), securing scores between 0.5 and 0.7. This confirms the Converge biobetter sits firmly in the 'clean' developability regime devoid of non-specific binding flags.
Antibody | BVP Score | dsDNA Score | Insulin Score |
Converge | 0.58 | 0.65 | 0.58 |
Cetuximab | 0.60 | 0.64 | 0.54 |
Cradle | 0.97 | 1.18 | 0.83 |
Infliximab (negative) | 0.57 | 0.77 | 0.53 |
Bevacizumab (medium) | 4.61 | 6.36 | 5.57 |
Bococizumab (high) | 84.57 | 100.17 | 93.19 |
Gantenerumab (high; reference) | 100 | 100 | 100 |
Table 5. PSR ELISA results for the three test antibodies and the calibration controls. MFI, median fluorescence intensity at 565 nm; S:B, sample-to-blank ratio; Score, PSR Score (% of Gantenerumab S:B). Lower values indicate lower polyspecificity.
Conclusion
In this work, we demonstrate that ConvergeAB™ can be applied in a fully zero-shot setting to generate a biobetter antibody with meaningfully improved properties over a clinically established therapeutic. Starting from cetuximab, the platform produced candidates that translated from in silico predictions to experimentally validated gains in binding kinetics, ultimately yielding an antibody with substantially enhanced affinity to EGFR. Importantly, this performance upgrade comes without a developability cost. Orthogonal measurements of hydrophobicity, particle size, thermal stability, and non-specific binding establish that the Converge variant matches or improves on cetuximab's biophysical profile. These results highlight the ability of ConvergeAB™ to efficiently navigate the antibody design space and its potential to accelerate the development of next-generation biologics with improved therapeutic performance.